Automating the Search for Artificial Life with Foundation Models
Abstract
With the recent Nobel Prize awarded for radical advances in protein discovery, foundation models (FMs) for exploring large combinatorial spaces promise to revolutionize many scientific fields. Artificial Life (ALife) has not yet integrated FMs, thus presenting a major opportunity for the field to alleviate the historical burden of relying chiefly on manual design and trial-and-error to discover the configurations of lifelike simulations. This paper presents, for the first time, a successful realization of this opportunity using vision-language FMs. The proposed approach, called NeuroLife, (1) finds simulations that produce target phenomena, (2) discovers simulations that generate temporally open-ended novelty, and (3) illuminates an entire space of interestingly diverse simulations. Because of the generality of FMs, NeuroLife works effectively across a diverse range of ALife substrates including Boids, Particle Life, Game of Life, Lenia, and Neural Cellular Automata. A major result highlighting the potential of this technique is the discovery of previously unseen Lenia and Boids lifeforms, as well as cellular automata that are open-ended like Conway's Game of Life. Additionally, the use of FMs allows for the quantification of previously qualitative phenomena in a human-aligned way. This new paradigm promises to accelerate ALife research beyond what is possible through human ingenuity alone.
Introduction
A core philosophy driving Artificial Life (ALife) is to study not only “life as we know it” but also “life as it could be”. Because ALife primarily studies life through computational simulations, this approach necessarily means searching through and mapping out an entire space of possible simulations rather than investigating any single simulation. By doing so, researchers can study why and how different simulation configurations give rise to distinct emergent behaviors. In this paper, we aim, for the first time, to automate this search through simulations with help from foundation models from AI.
While the specific mechanisms for evolution and learning within ALife simulations are rich and diverse, a major obstacle so far to fundamental advances in the field has been the lack of a systematic method for searching through all the possible simulation configurations themselves. Without such a method, researchers must resort to intuitions and hunches when devising perhaps the most important aspect of an artificial world—the rules of the world itself.
Part of the challenge is that large-scale interactions of simple parts can lead to complex emergent phenomena that are difficult, if not impossible, to predict in advance. This disconnect between the simulation configuration and its resulting behavior makes it difficult to intuitively design simulations that exhibit self-replication, ecosystem-like dynamics, or open-ended properties. As a result, the field often delivers manually designed simulations tailored to simple and anticipated outcomes, limiting the potential for unexpected discoveries.
Given this present improvisational state of the field, a method to automate the search for simulations themselves would transform the practice of ALife by significantly scaling the scope of exploration. Instead of probing for rules and interactions that feel right, researchers could refocus their attention to the higher-level question of how to best describe the phenomena we ultimately want to emerge as an outcome, and let the automated process of searching for those outcomes then take its course.
Describing target phenomena for simulations is challenging in its own right, which in part explains why automated search for the right simulation to obtain target phenomena has languished. Of course, there have been many previous attempts to quantify ALife through intricate measures of life, complexity, or “interestingness”. However, these metrics almost always fail to fully capture the nuanced human notions they try to measure.
While we don't yet understand why or how our universe came to be so complex, rich, and interesting, we can still use it as a guide to create compelling ALife worlds. Foundation models (FMs) trained on large amounts of natural data possess representations often similar to humans and may even be converging toward a “platonic” representation of the statistics of our real world. This novel property makes them appealing candidates for quantifying human notions of complexity in ALife.
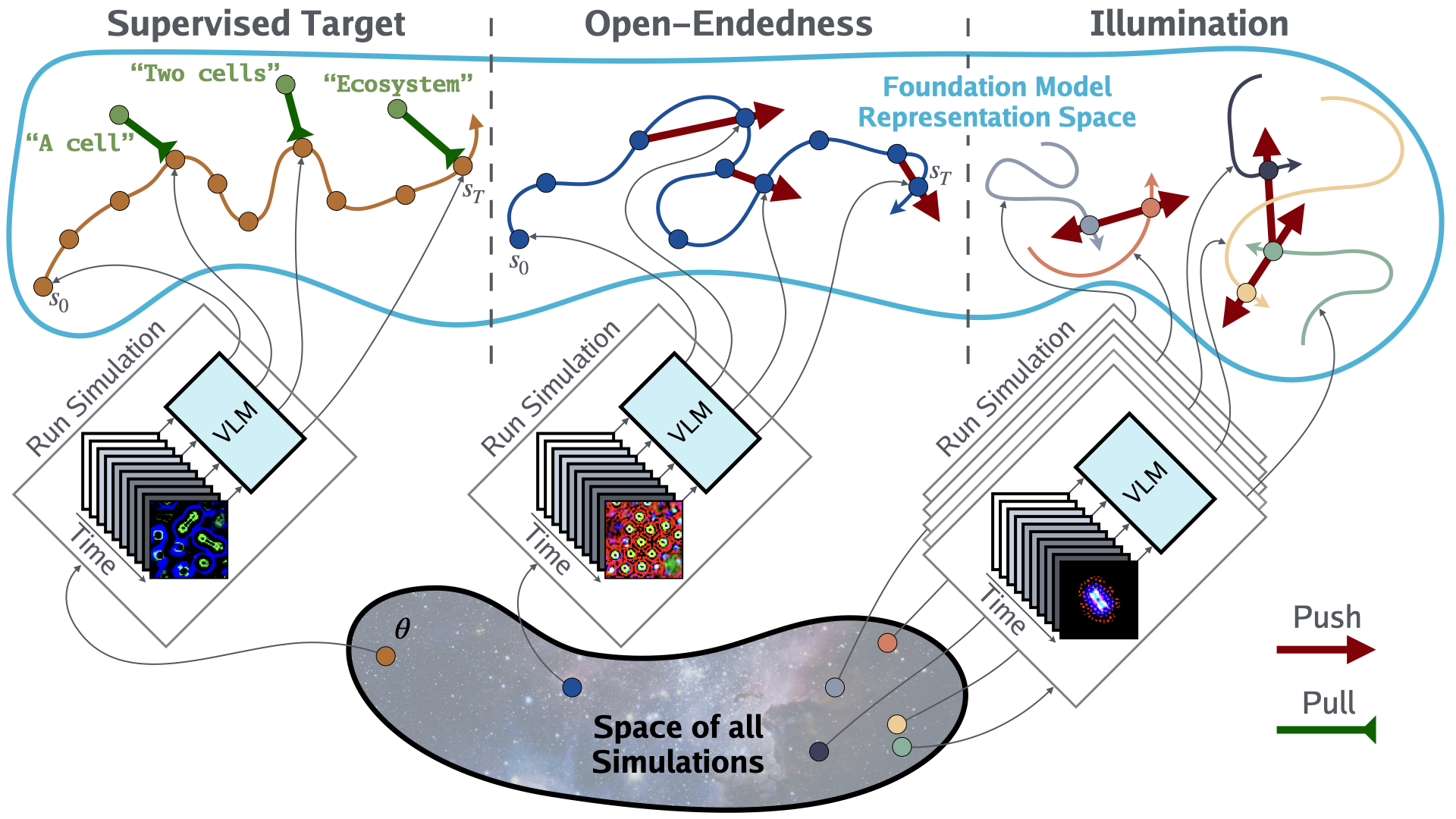
In this spirit, we propose a new paradigm for ALife research called NeuroLife. The researcher starts by defining a set of simulations of interest, referred to as the substrate. Then, as shown in the figure above, NeuroLife enables three distinct methods for FMs to identify interesting ALife simulations: (1) Supervised Target—Searching for a simulation that produces a specified target event or sequence of events, facilitating the discovery of arbitrary worlds or those similar to our own. (2) Open-Endedness—Searching for a simulation that produces temporally open-ended novelty in the FM representation space, thereby discovering worlds that are persistently interesting to a human observer. (3) Illumination—Searching for a set of interestingly diverse simulations, enabling the illumination of alien worlds.
The promise of this new automated approach is demonstrated on a diverse range of ALife substrates including Boids, Particle Life, Game of Life, Lenia, and Neural Cellular Automatas. In each substrate, NeuroLife discovered previously unseen lifeforms and expanded the frontier of emergent structures in ALife. For example, NeuroLife revealed exotic flocking patterns in Boids, new self-organizing cells in Lenia, and identified cellular automata which are open-ended like the famous Conway's Game of Life. In addition to facilitating discovery, NeuroLife's FM framework allows for quantitative analysis of previously qualitative phenomena in ALife simulations, providing a human-like approach to measuring complexity. NeuroLife is agnostic to both the specific FM and the simulation substrate, enabling compatibility with future FMs and ALife substrates.
Overall, our new FM-based paradigm serves as a valuable tool for future ALife research by stepping towards the field's ultimate goal of exploring the vast space of artificial life forms. To the best of our knowledge, this is the first work to drive ALife simulation discovery through foundation models.
Methods: Automated Search for Artificial Life
An ALife substrate encompasses any set of ALife simulations of interest (e.g. the set of all Lenia simulations). These could vary in the initial states, transition rules, or both. The substrate is parameterized by θ, which defines a single simulation with three components: the initial state distribution, the forward dynamics step function, and the rendering function, which transforms the state into an image.
While parameterizing and searching for a renderer is often not needed, it becomes necessary when dealing with state values that are uninterpretable a priori. Chaining these terms together, we define a function of θ that samples an initial state, runs the simulation for T steps, and renders that final state as an image. Finally, two additional functions embed images and natural language text through the vision-language FM, along with a corresponding inner product to facilitate similarity measurements for that embedding space.
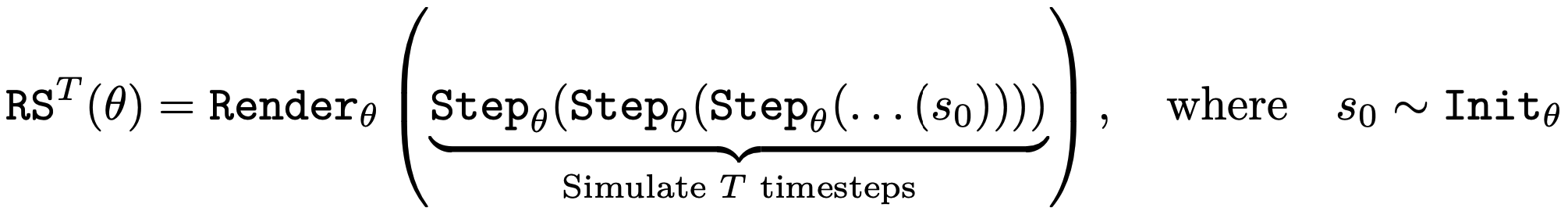
Supervised Target
An important goal in ALife is to find simulations where a desired event or sequence of events take place. Such discovery would allow researchers to identify worlds similar to our own or test whether certain counterfactual evolutionary trajectories are even possible in the given substrate, thus giving insights about the feasibility of certain lifeforms. For this purpose, NeuroLife searches for a simulation that produces images that match a target natural language prompt in the FM's representation. The researcher has control of which prompt, if any, to apply at each timestep.
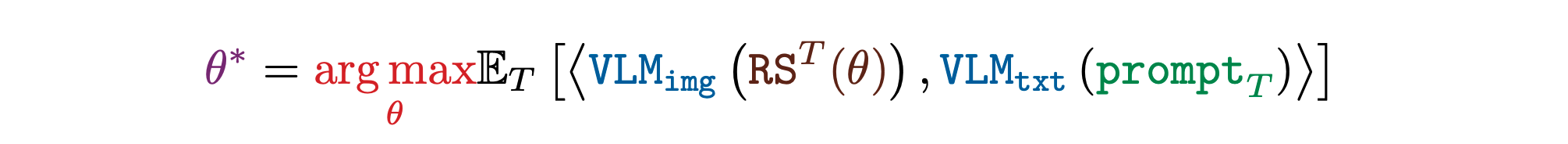
Open-Endedness
A grand challenge of ALife is finding open-ended simulations. Finding such worlds is necessary for replicating the explosion of never-ending interesting novelty that the real world is known for. Although open-endedness is subjective and hard to define, novelty in the right representation space captures a general notion of open-endedness. This formulation outsources the subjectivity of measuring open-endedness to the construction of the representation function, which embodies the observer. In this paper, the vision-language FM representations act as a proxy for a human's representation.
With this novel capability, NeuroLife searches for a simulation which produces images that are historically novel in the FM's representation. Some preliminary experiments showed that historical nearest neighbor novelty produces better results than variance based novelty.
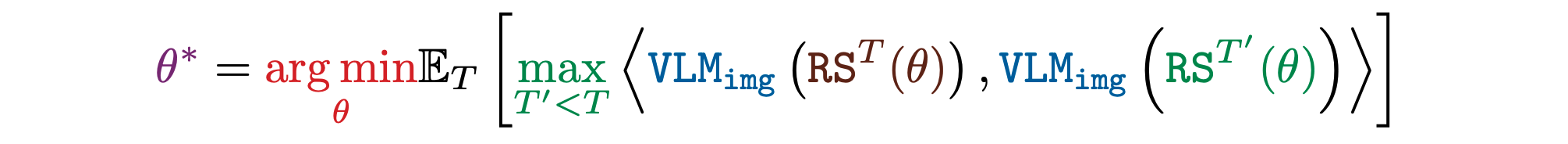
Illumination
Another key goal in ALife is to automatically illuminate the entire space of diverse phenomena that can emerge within a substrate, motivated by the quest to understand “life as it could be”. Such illumination is the first step to mapping out and taxonomizing an entire substrate. Towards this aim, NeuroLife searches for a set of simulations that produce images that are far from their nearest neighbor in the FM's representation. We find that nearest neighbor diversity produces better illumination than variance based diversity.
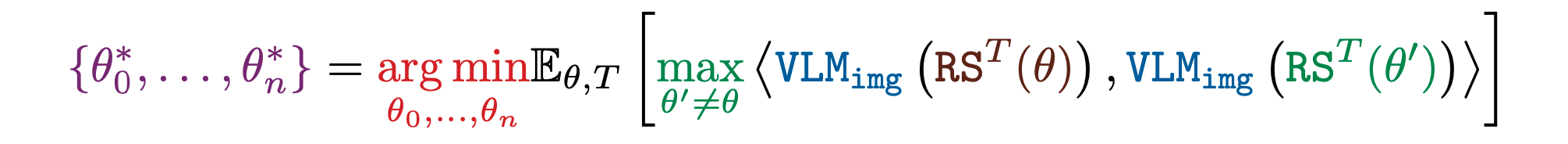
Experiments
This section experimentally validates the effectiveness of NeuroLife across various substrates, then presents novel quantitative analyses of some of the discovered simulations, facilitated by the FM. Before presenting the experiments, here is a summary of the FMs and substrates used. The appendix includes additional details about the substrates and experimental setups, as well as supplementary experiments.
Searching for Target Simulations
The effectiveness of searching for target simulations specified by a single prompt is explored in Lenia, Boids, and Particle Life. The supervised target equation is optimized with the prompt applied once after T simulation timesteps. CLIP is the FM and Sep-CMA-ES is the optimization algorithm. The figure below shows the optimization works well from a qualitative perspective at finding simulations matching the specified prompt. Some of the failure modes suggest that when optimization fails, it is often caused by the lack of expressivity of the substrate rather than the optimization process itself.
We investigate the effectiveness of searching for simulations producing a target sequence of events using the NCA substrate. We optimize the supervised target equation with a list of prompts, each applied at evenly spaced time intervals of the simulation rollout. We use CLIP for the FM. Following the original NCA paper, we use backpropagation through time and gradient descent with the Adam optimizer for the optimization algorithm. The figure below shows it is possible to find simulations that produce trajectories following a sequence of prompts. By specifying the desired evolutionary trajectories and employing a constraining substrate, NeuroLife can identify update rules that embody the essence of the desired evolutionary process. For instance, when the sequence of prompts is “one cell” then “two cells”, the corresponding update rule inherently enables self-replication.
Searching for Open-Ended Simulations
To investigate the effectiveness of searching for open-ended simulations, we use the Life-Like CAs substrate and optimize the open-endedness score. CLIP serves as the FM. Because the search space is relatively small with only 262,144 simulations, brute force search is employed. The figure below reveals the potential for open-endedness in the Life-like CAs. The famous Conway's Game of Life ranks among the top 5% most open-ended CAs according to our open-endedness metric. The top subfigure shows the most open-ended CAs demonstrate nontrivial dynamic patterns that lie on edge of chaos, since they neither plateau or explode. The bottom left subfigure traces out the trajectories of three CAs in CLIP space over simulation time. The bottom right subfigure visualizes all Life-Like CAs with a UMAP plot of their CLIP embeddings colored by open-endedness score, and shows that meaningful structure emerges: the most open-ended CAs lie close together on a small island outside the main island of simulations.
Illuminating Entire Substrates
We use the Lenia and Boids substrates to study the effectiveness of the illumination algorithm. CLIP is the FM. A custom genetic algorithm performs the search: at each generation, it randomly selects parents, creates mutated children, then keeps the most diverse subset of solutions. The resultant set of simulations is shown in the “Simulation Atlas” in the figure below. This visualization highlights the diversity of the discovered behaviors organized by visual similarity. In Lenia, NeuroLife discovers many previously unseen lifeforms resembling cells and bacteria organized by color and shape. In Boids, NeuroLife rediscovers flocking behavior, as well as additional behaviors such as snaking, grouping, circling, and other variations. Larger simulation atlases are shown in appendix.
Quantifying ALife
Not only can FMs facilitate the search for interesting phenomena, but they also enable the quantification of phenomena previously only amenable to qualitative analysis, as shown in this section. The following figures show different ways of quantifying the emergent behaviors of these complex systems.
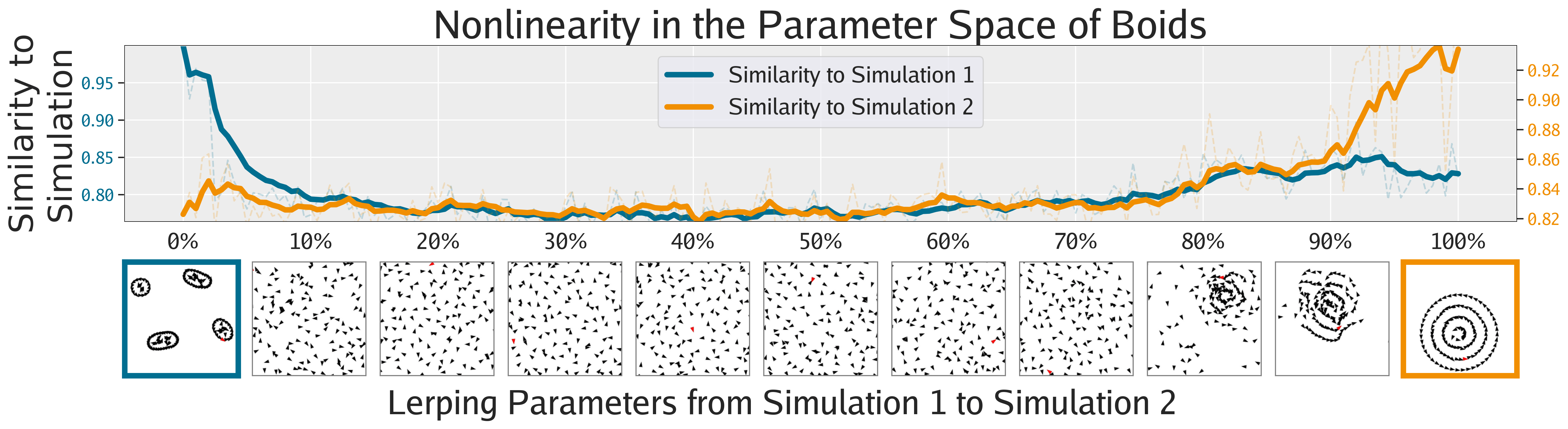
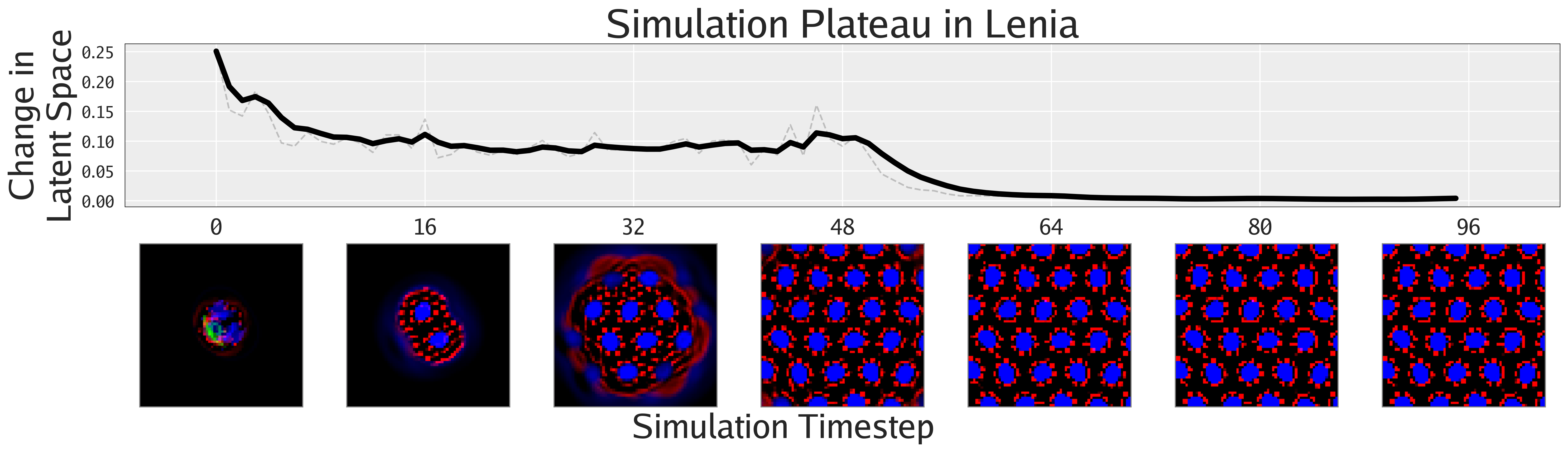
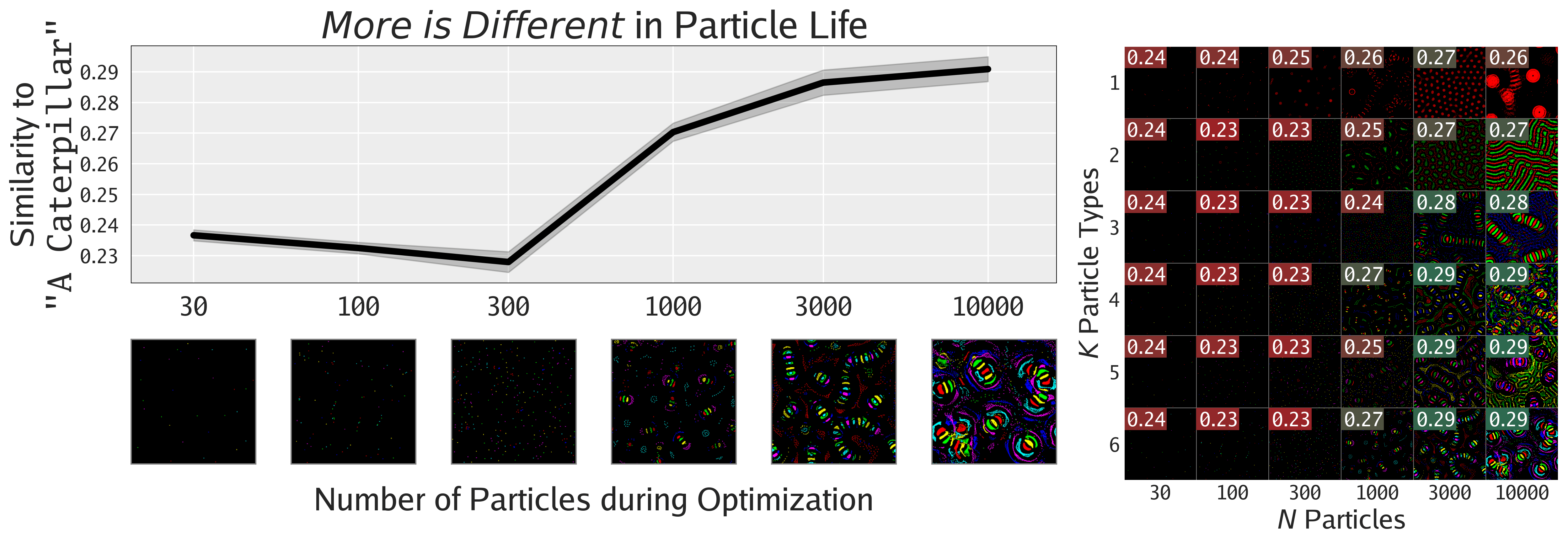
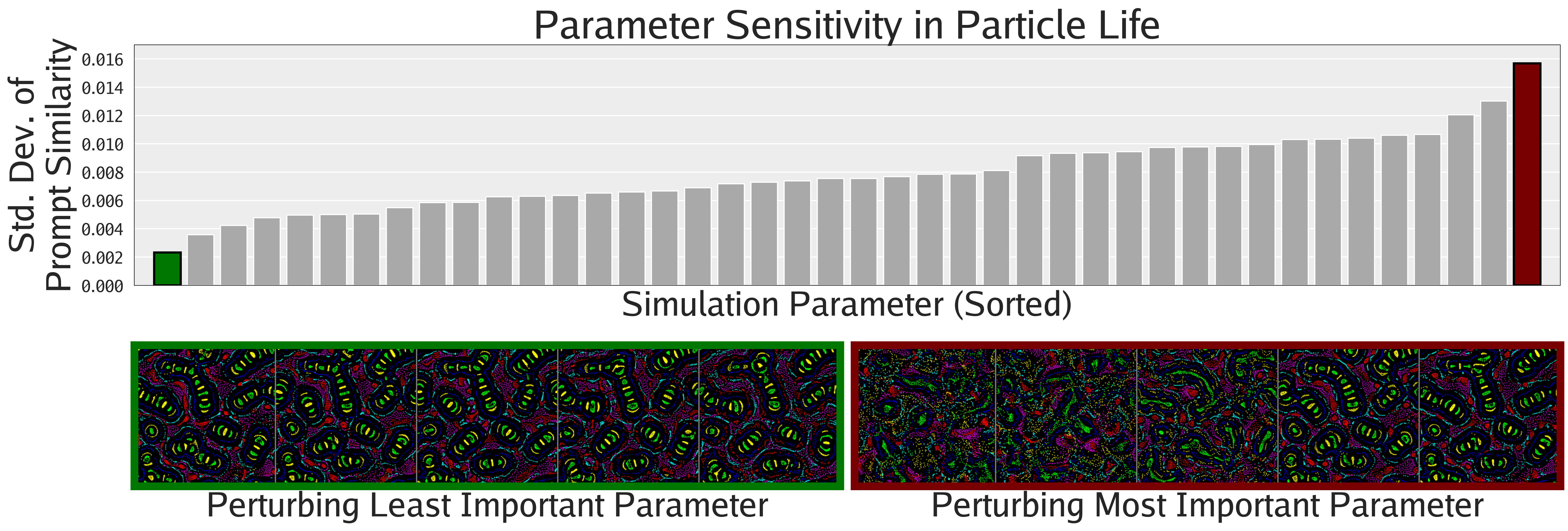
Agnostic to Foundation Model
To study the importance of using the proper representation space, we ablate the FM used during illumination of Lenia and Boids. We use CLIP, DINOv2, and a low level pixel representation. As shown in the figure below, for producing human-aligned diversity, CLIP seems slightly better than DINOv2, but both are qualitatively better than the pixel representation. This result highlights the importance of deep FM representations over low-level metrics when measuring human notions of diversity.
Related Works
ALife is a diverse field that studies life through artificial simulations, with the key difference from biology being its pursuit of the general properties of all life rather than just the specific instantiation of life on Earth. ALife systems range widely from cellular automata to neural network agents, but the field generally focuses on emergent phenomena like self-organization, open-ended evolution, agentic behavior, and collective intelligence. These ALife ideas have also trickled into AI.
Many substrates are used in ALife to study phenomena at different levels of abstraction and imposed structure, ranging from modeling chemistry to societies. Conway's Game of Life and other “Life-Like” cellular automatas (CA) were critical to the field in the early days and are used to study how complexity may emerge from simple local rules. Lenia generalizes these to continuous dynamics, and inspired future variants like FlowLenia and ParticleLenia. Neural Cellular Automata (NCA) further generalize Lenia by parameterizing the update rule with a neural network. Instead of operating in a 2-D grid, ParticleLife (or Clusters) uses particles in euclidean space interacting with each other to create dynamic self-organizing patterns. Similarly, Boids uses bird-like objects to model the flocking behavior of real birds and fish.
Automatic search has been a useful tool in ALife whenever the target outcome is well defined. In the early days, genetic algorithms were used to evolve CAs to produce a target computation. Novelty search is a search algorithm inspired by ALife but requires a good representation space to be effective. MAP-Elites is a search algorithm which searches along two predefined axis of interest. Intrinsically motivated discovery uses search in the representation space of an autoencoder to discover new self-organizing patterns. LeniaBreeder uses MAP-Elites to search for organisms which have specific properties for e.g. mass and speed.
Many attempts have been made to quantify complexity. In information theory, Kolmogorov complexity measures the length of the shortest computer program that produces an artifact. Rather than measuring the complexity of an artifact directly, sophistication measures the complexity of the set in which the artifact is a “generic” member. Stemming from biochemistry, assembly theory hopes to quantify evolution by measuring the minimal number of steps required to assemble an artifact from atomic building blocks or previously assembled pieces. Although theoretically compelling, these metrics are not computable or fail to capture the nuanced human notions of complexity.
Open-endedness (OE) is the ability of a system to keep generating interesting artifacts forever, which is one of the defining features of natural evolution. Many necessary conditions for OE have been identified but are yet to be realized. There have been some attempts at quantifying OE, but some argue that OE cannot be quantified by definition. In one case study, human intervention was essential for achieving OE evolution, suggesting that OE may depend on novelty within a particular representation space.
Large pretrained neural networks, often referred to as foundation models (FMs), are currently revolutionizing many scientific domains. In medicine, FMs transformed the drug discovery process by enabling accurate predictions of protein folding. In robotics, LLMs have automated the design of reward functions. In physics, large models are used to predict complex systems and are later distilled into symbolic equations. In this work, we use CLIP, an image-language embedding model trained with contrastive learning to align text-image representations on an internet-scale dataset. CLIP's simplicity and generality enable it to effectively guide generation, and we apply CLIP to search for ALife simulations instead of static images, resulting in analyzable artifacts that provide valuable insights for ALife research.
Conclusion
This project launches a new paradigm in ALife by taking the first step towards using FMs to automate the search for interesting simulations. Our approach is effective in finding target, open-ended, and diverse simulations over a wide spectrum of substrates. Additionally, FMs enable the quantification of many qualitative phenomena in ALife, offering a path to replacing low-level complexity metrics with deep representations aligned with humans.
Because this project is agnostic to the FM and substrate used, it raises the question of which ones to use. The choice of FMs seems to not matter much from our experiments, and FMs in general may also be converging to similar representations of reality. The proper substrate largely depends on the phenomena that is being studied (e.g. self-organization, open-ended evolution, etc.). The most expressive substrate would simply parameterize all the RGB pixels of an entire video, but is useless for studying emergence. The most insightful substrates bake in as little information as possible, while maintaining vast emergent capabilities.
Eventually, with the proper substrate, more powerful FMs, and enough compute, this paradigm may allow researchers to automatically search for worlds which start off as “simple cells in primordial soup”, then undergo “a Cambrian explosion of complexity”, and eventually become “an artificial alien civilization”. Researchers could alternatively search for hypothetical worlds where life evolves without DNA. Finding open-ended worlds would solve one of ALife's grand challenges. Illuminating such a substrate could help map the space of possible lifeforms and intelligences, giving a taxonomy of life as it could be in the computational universe.
This work can be generalized by replacing the image-language FM with video-language FMs that natively process the temporal nature of simulations or with 3-D FMs to handle 3-D simulations. To leverage the recent advances of LLMs, images can be converted to text via image-to-text models, allowing all analyses to be done in text space. Instead of searching for ALife simulations, a similar approach could be constructed for low-level physics research. At a meta-level, LLMs could be useful for generating code that describes the substrates themselves, driven by higher-level research agendas.
Acknowledgments
This work was supported by an NSF GRFP Fellowship and by ONR MURI grant N00014-22-1-2740.
Citation
If you find this work useful, please cite it as follows: